Mechanisms of Genetic Control and Sequence Variation
Regulation of Gene Expression
Gene regulation allows cells to control which genes are turned on (expressed) or turned off (repressed) at any given time. This conservation of energy and space is vital for survival. While the central dogma ($DNA \rightarrow RNA \rightarrow Protein$) applies to all life, the methods of regulation differ significantly between prokaryotes and eukaryotes.
Prokaryotic Gene Regulation: The Operon Model
Prokaryotes (bacteria) often organize genes with related functions into groups called operons. These genes are transcribed together into a single mRNA molecule, allowing protein synthesis to be controlled as a single unit.
Components of an Operon
- Promoter: A specific DNA sequence where RNA Polymerase attaches to begin transcription.
- Operator: A DNA sequence located between the promoter and the structural genes. It acts as the "on/off switch."
- Structural Genes: The genes coding for the metabolic enzymes.
- Regulatory Gene: A gene located upstream (outside the operon) that codes for a repressor protein. This protein can bind to the operator to block transcription.
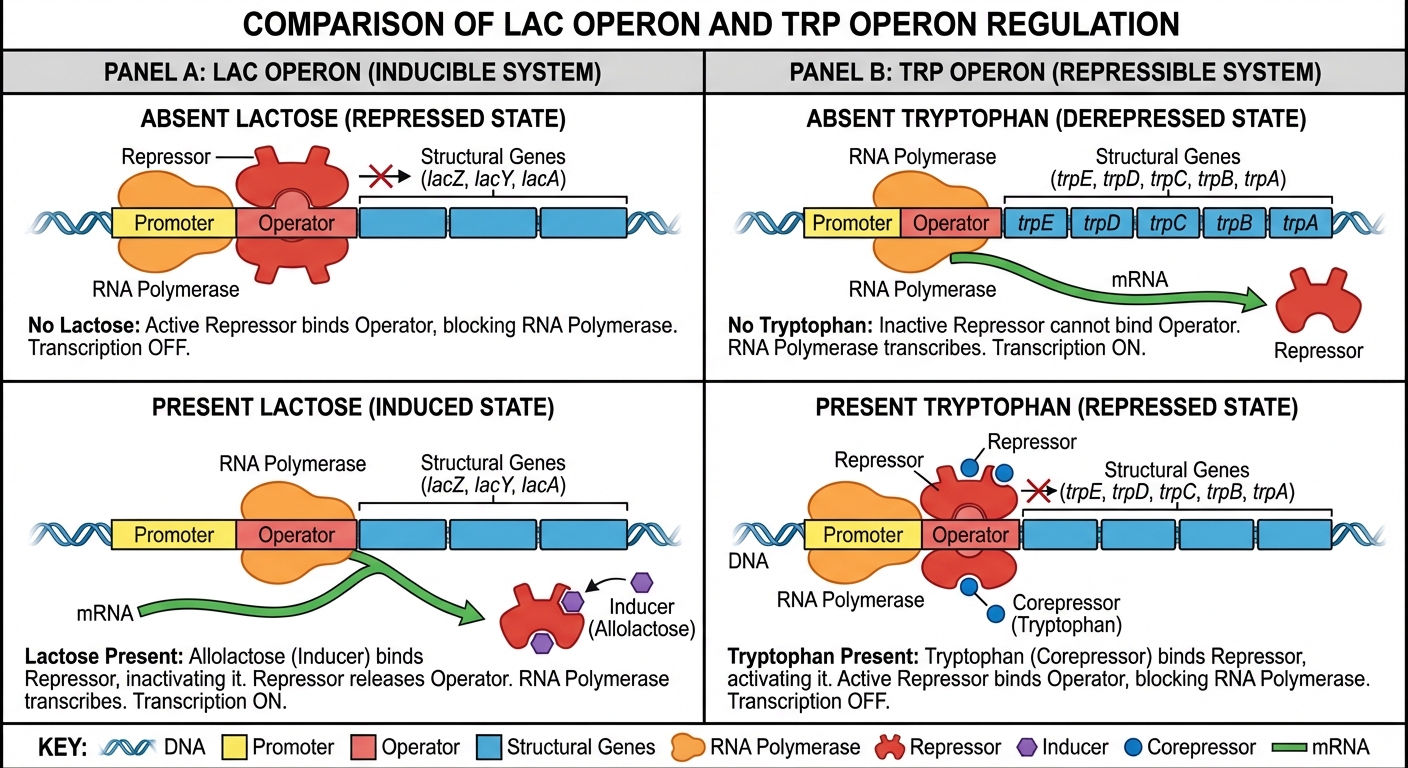
Inducible vs. Repressible Operons
There are two main types of negative gene regulation in prokaryotes:
1. Inducible Operons (Example: lac operon)
- Default State: OFF.
- Function: Usually limits the production of enzymes that break down (catabolize) specific substances until that substance is present.
- Mechanism:
- The repressor is normally active and bound to the operator, blocking RNA polymerase.
- When the inducer (allolactose, an isomer of lactose) binds to the repressor, the repressor changes shape and detaches from the operator.
- Transcription turns ON.
2. Repressible Operons (Example: trp operon)
- Default State: ON.
- Function: Usually controls the synthesis of organic molecules (anabolism) required by the cell (like amino acids).
- Mechanism:
- The repressor is normally inactive, allowing transcription to proceed.
- When the corepressor (tryptophan) accumulates, it binds to the repressor, activating it.
- The active repressor binds to the operator, helping to turn transcription OFF to prevent overproduction.
Eukaryotic Gene Regulation
Eukaryotes do not use operons. Because eukaryotic cells have a nucleus and complex organelles, regulation can occur at many stages: chromatin modification, transcription, RNA processing, translation, and protein degradation.
1. Chromatin Structure and Epigenetics
The physical packing of DNA affects gene availability.
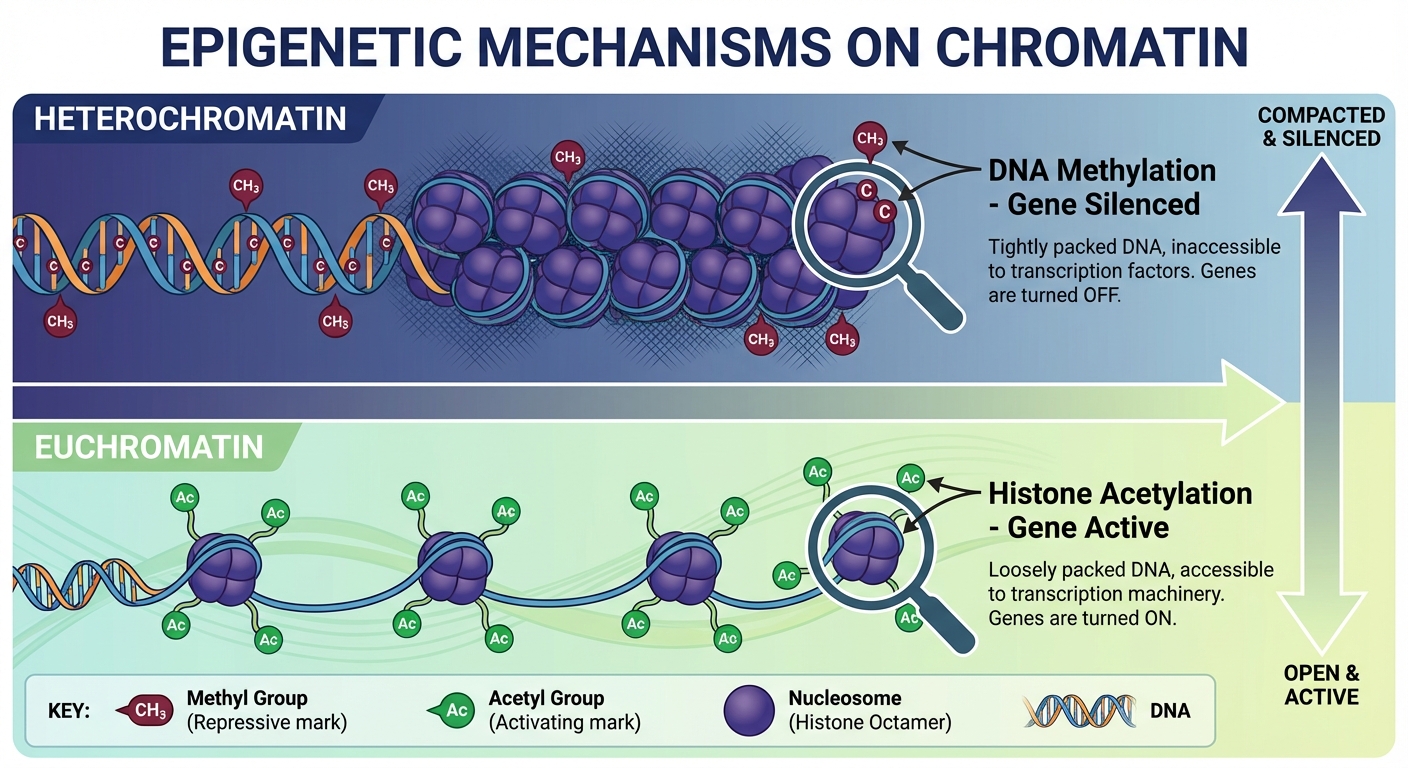
- Histone Acetylation: The attachment of acetyl groups ($-COCH_3$) to histone tails. This loosens chromatin structure, making DNA transcriptionally active.
- DNA Methylation: The addition of methyl groups ($-CH_3$) to DNA bases (usually cytosine). This condenses, or causes the DNA to coil more tightly, rendering the gene inactive.
Epigenetics refers to heritable changes in gene expression that do not involve changes to the underlying DNA sequence.
2. Transcriptional Control
To initiate transcription, eukaryotic RNA polymerase requires the assistance of proteins called Transcription Factors.
- General Transcription Factors: Bind to the promoter (often at a specific sequence called the TATA box) to help RNA polymerase bind.
- Specific Transcription Factors (Activators/Repressors): Bind to control elements called Enhancers (far upstream) or Silencers. Interaction between enhancers and the promoter is mediated by DNA bending.
3. Alternative RNA Splicing
This is a crucial mechanism for efficiency. During RNA processing (in the nucleus), different mRNA molecules are produced from the same primary transcript depending on which RNA segments are treated as exons (coding) and which as introns (non-coding).
- Result: One gene can code for multiple distinct proteins.
Gene Expression and Cell Specialization
All somatic cells in a multicellular organism contain the exact same genome. However, a neuron looks and functions differently from a liver cell. This differentiation is due to Differential Gene Expression.
Determinants of Cell Specialization
- Cytoplasmic Determinants: Maternal substances in the egg that influence early development. Basically, the uneven distribution of RNA/proteins in positions of the fertilized egg leads to different gene expression in daughter cells.
- Induction: Signal molecules from embryonic cells cause transcriptional changes in nearby target cells (cell-cell communication).
Small Regulatory RNAs
Non-coding RNAs play a role in silencing genes after transcription:
- MicroRNAs (miRNAs) and Small Interfering RNAs (siRNAs) can bind to mRNA and degrade it or block translation, effectively silencing the gene.
Mutations
A Mutation is a permanent change in the nucleotide sequence of the genome. While often viewed negatively, mutations are the primary source of genetic variation, which is the raw material for evolution.
Types of Gene Mutations
1. Point Mutations
A change in a single nucleotide pair of a gene.
| Type | Description | Consequence |
|---|---|---|
| Silent | A nucleotide substitution that codes for the same amino acid. | No change in protein function (due to redundancy in the genetic code). |
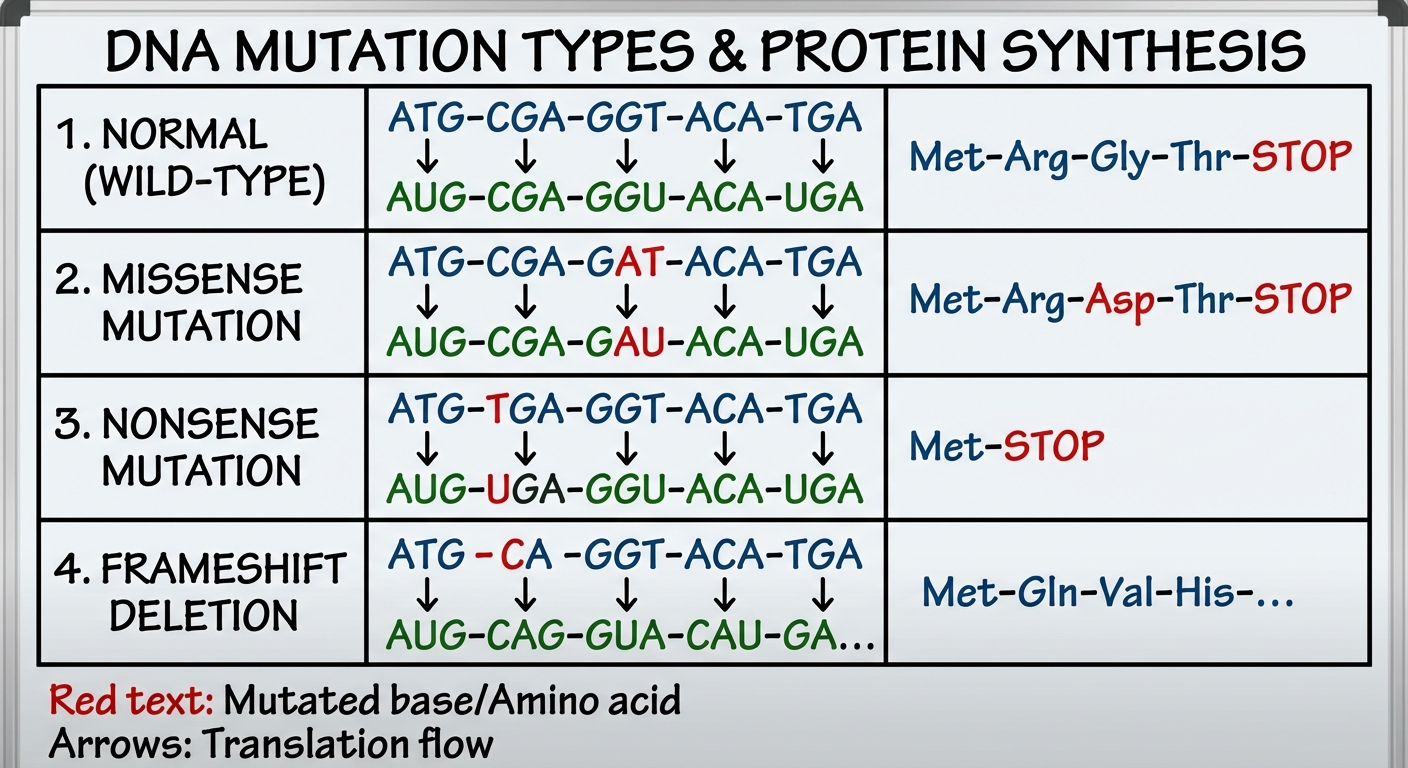
| Missense | A substitution that codes for a different amino acid. | Can range from little effect to severe loss of function (e.g., Sickle Cell Anemia). |
| Nonsense | A substitution that changes an amino acid codon into a STOP codon. | Leads to a truncated (shortened), usually non-functional protein. |
2. Frameshift Mutations
Caused by Insertions or Deletions of nucleotides that are not in multiples of three.
- Because the mRNA is read in triplets (codons), acting as a "reading frame," adding or removing one base shifts the grouping of every subsequent beat.
- Result: Extensive missense immediately causing the wrong amino acids to be translated, usually followed by an immediate nonsense (premature stop). These are almost always disastrous for the protein.
Chromosomal Mutations
Large-scale changes affecting whole chromosomes or large segments:
- Duplication: A segment is repeated.
- Deletion: A segment is removed.
- Inversion: A segment is reversed within the chromosome.
- Translocation: A segment moves from one chromosome to a non-homologous chromosome.
Causes and Effects
Causes
- Spontaneous: Errors in DNA replication, recombination, or repair.
- Mutagens: Physical or chemical agents (e.g., X-rays, UV light, carcinogens) that interact with DNA.
Evolutionary Context
It is critical to remember that mutations are random. An organism cannot "develop" a mutation because it needs it.
- Neutral: Most mutations have no effect or are in non-coding regions.
- Detrimental: Changes protein shape in a way that hinders function.
- Beneficial: Occasionally, a mutation confers a survival advantage (e.g., antibiotic resistance in bacteria), which drives natural selection.
Common Mistakes & Pitfalls
- Operon Confusion: Students often confuse repressible with inducible.
- Tip: Remember LOCK and KEY. In the lac operon (inducible), the lock is ON, and you need the key (lactose) to open it. in the trp operon (repressible), the safety is OFF, and you use the key (tryptophan) to lock it.
- Gene Regulation $\neq$ DNA Replication: Do not confuse transcription factors (Unit 6) with DNA Polymerase/Ligase (Unit 5). Regulation is about reading the DNA, not copying it.
- Substitution vs. Frameshift: Students often underestimate frameshifts. A point mutation changes one letter; a frameshift changes the entire sentence from that point forward.
- All Mutations are Bad: Avoid this absolute language. Positive mutations are rare but essential for evolution.
- Genomic Equivalence: Don't forget that your eye cells have the genes to make stomach acid—they just have those genes turned off (methylated).
- Epigenetic Mnemonic:
- Methylation = Mute (Turns off).
- Acetylation = Ace / Active (Turns on).