Genetics: The Molecular Basis of Inheritance
Unit 6: Gene Expression and Regulation
Structure of DNA and RNA
The fundamental unit of life's genetic code is the nucleotide. Understanding the structural differences between DNA and RNA is crucial for mastering gene expression.
Nucleotide Structure
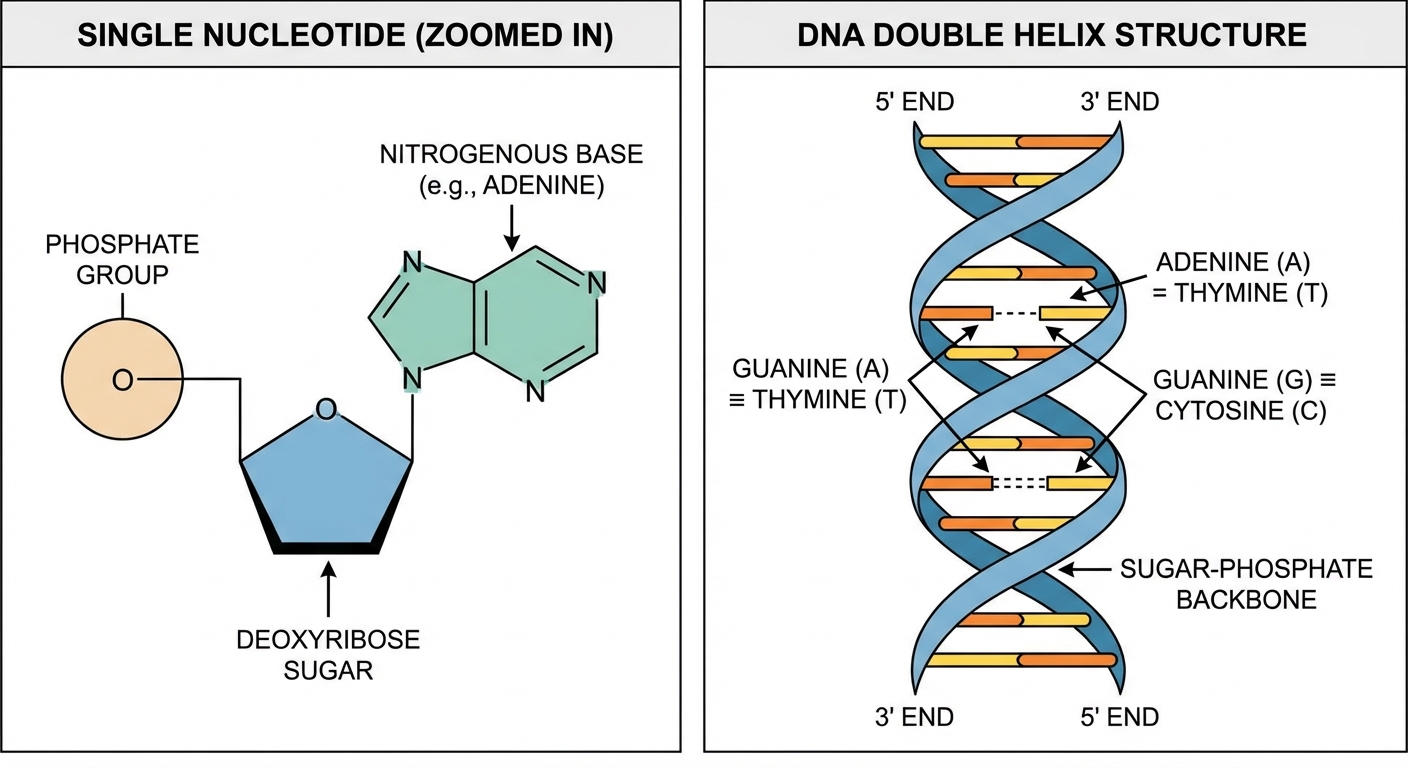
A nucleotide consists of three parts linked together:
- A Five-Carbon Sugar:
- Deoxyribose in DNA (lacks an oxygen at the 2' carbon).
- Ribose in RNA (has an -OH group at the 2' carbon).
- A Phosphate Group: Attached to the 5' carbon of the sugar. This gives DNA its negative charge.
- A Nitrogenous Base: Attached to the 1' carbon.
Nitrogenous Bases
Bases are categorized by their chemical structure:
- Purines: Double-ring structures. Mnemonic: "Pure As Gold".
- Adenine
- Guanine
- Pyrimidines: Single-ring structures. Mnemonic: "Cut The Py".
- Cytosine
- Thymine (DNA only)
- Uracil (RNA only)
The Double Helix and Base Pairing
DNA exists as a double helix of two complementary strands running antiparallel to each other.
Directionality: One strand runs while the opposite runs .
- The 5' end terminates with a Phosphate group.
- The 3' end terminates with a Hydroxyl (-OH) group.
- Crucial Rule: New nucleotides can ONLY be added to the 3' end.
Chargaff's Rules (Base Pairing):
- A pairs with T (linked by 2 Hydrogen Bonds).
- G pairs with C (linked by 3 Hydrogen Bonds).
- Note: The G-C bond is stronger due to the extra hydrogen bond, making G-C rich DNA harder to separate.
Organizing the Genome
- Prokaryotes: typically have a single, circular chromosome found in the nucleoid region, plus small rings of DNA called plasmids.
- Eukaryotes: have multiple linear chromosomes housed in a nucleus.
- Histones: Positively charged proteins that DNA wraps around.
- Nucleosomes: The "beads on a string" structure formed by DNA wrapping around histones.
- Euchromatin: Loosely packed DNA; active and available for transcription.
- Heterochromatin: Tightly packed, condensed DNA; generally inactive.
DNA Replication
Replication is semiconservative: each new DNA molecule consists of one original parental strand and one newly synthesized strand.
Key Enzymes in Replication
| Enzyme | Function |
|---|---|
| Helicase | Unzips the double helix by breaking hydrogen bonds, creating a replication fork. |
| Topoisomerase | Relieves the strain/supercoiling ahead of the replication fork by cutting and rejoining DNA strands. |
| Primase | Synthesizes a short RNA primer to provide a 3' OH group for DNA polymerase to start. |
| DNA Polymerase III | Adds DNA nucleotides to the 3' end of the growing strand. |
| DNA Polymerase I | Removes RNA primers and replaces them with DNA. |
| Ligase | "Glues" DNA fragments (Okazaki fragments) together by forming phosphodiester bonds. |
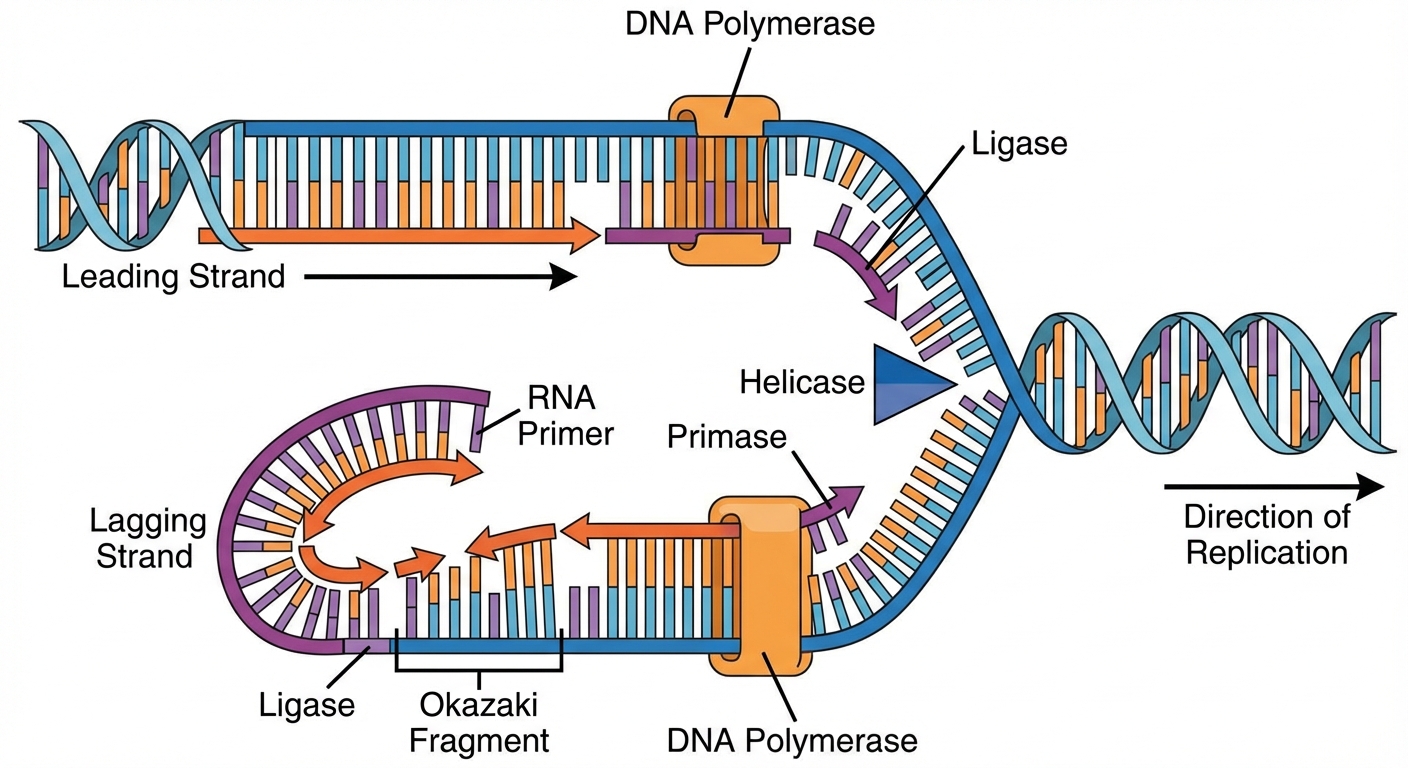
The Process
Replication begins at origins of replication. Because DNA Polymerase only works in the direction, the two strands are built differently:
- Leading Strand: Synthesized continuously toward the replication fork.
- Lagging Strand: Synthesized discontinuously away from the fork in short segments called Okazaki fragments.
The "End" Problem
In eukaryotes, the machinery cannot copy the very tips of linear chromosomes.
- Telomeres: Repetitive non-coding sequences at the ends of chromosomes that act as protective caps. They shorten with each division, contributing to aging.
Transcription: Copying DNA to RNA
This is the synthesis of RNA using a DNA template.
The Central Dogma
(Note: Retroviruses violate this by using Reverse Transcriptase to go RNA $\rightarrow$ DNA).
Steps of Transcription
- Initiation: RNA Polymerase binds to a region called the promoter (often contains the TATA box in eukaryotes). Transcription factors help mediate this binding.
- Elongation: RNA Polymerase unwinds DNA and adds RNA nucleotides to the 3' end of the growing RNA molecule.
- Template Strand (Antisense): The strand used as a guide. The RNA is complementary to this.
- Coding Strand (Sense): The non-template strand. The RNA sequence looks exactly like this strand (except U replaces T).
- Termination: The enzyme detaches after transcribing a terminator sequence.
RNA Processing (Eukaryotes Only)
Before leaving the nucleus, pre-mRNA must be modified:
- 5' GTP Cap: Added to the 5' end. Protects RNA and helps ribosome attachment.
- Poly-A Tail: Added to the 3' end. Increases stability and helps export from the nucleus.
- Splicing:
- Introns: Non-coding regions are cut out.
- Exons: Coding regions are spliced together (expressed).
- Alternative Splicing: A single gene can code for multiple proteins depending on which exons are kept.
Translation: RNA to Protein
Translation occurs at the ribosome (in the cytoplasm or on the rough ER).
Key Players
- mRNA: Carries the message.
- tRNA (Transfer RNA): The "translator." It has an anticodon on one end and a specific amino acid on the other.
- Ribosome (rRNA + protein): Has three sites: A (Aminoacyl), P (Peptidyl), and E (Exit).
The Genetic Code
- Codon: A 3-nucleotide sequence on mRNA.
- Redundancy: There are 64 codons but only 20 amino acids. Most amino acids are coded for by more than one codon (helps protect against mutations).
- Start Codon: AUG (Methionine).
- Stop Codons: UAA, UAG, UGA (signal termination).
The Process
- Initiation: The small ribosomal subunit binds to mRNA at the start codon (AUG). The tRNA carrying Methionine binds to the P-site.
- Elongation:
- Codon Recognition: Next tRNA enters the A-site.
- Peptide Bond Formation: The amino acid chain moves from the P-site tRNA to the A-site tRNA.
- Translocation: The ribosome shifts down one codon. Empty tRNA leaves via the E-site.
- Termination: A stop codon is reached. A release factor binds, dismantling the complex.
Common Mistake: Students often confuse the template strand with the coding strand. Remember: The mRNA is complementary to the Template, making it identical to the Coding strand (swapping U for T).
Regulation of Gene Expression
Cells must control which genes are expressed (turned on) to conserve energy and strictly control cell differentiation.
Prokaryotic Regulation: Operons
Bacteria group related genes under a single promoter, called an operon.
Repressible Operon (e.g., trp operon):
- Default State: ON.
- Function: Anabolic (building things).
- Mechanism: When Tryptophan is present (corepressor), it binds to the repressor, activating it. The repressor binds to the operator, stopping transcription.
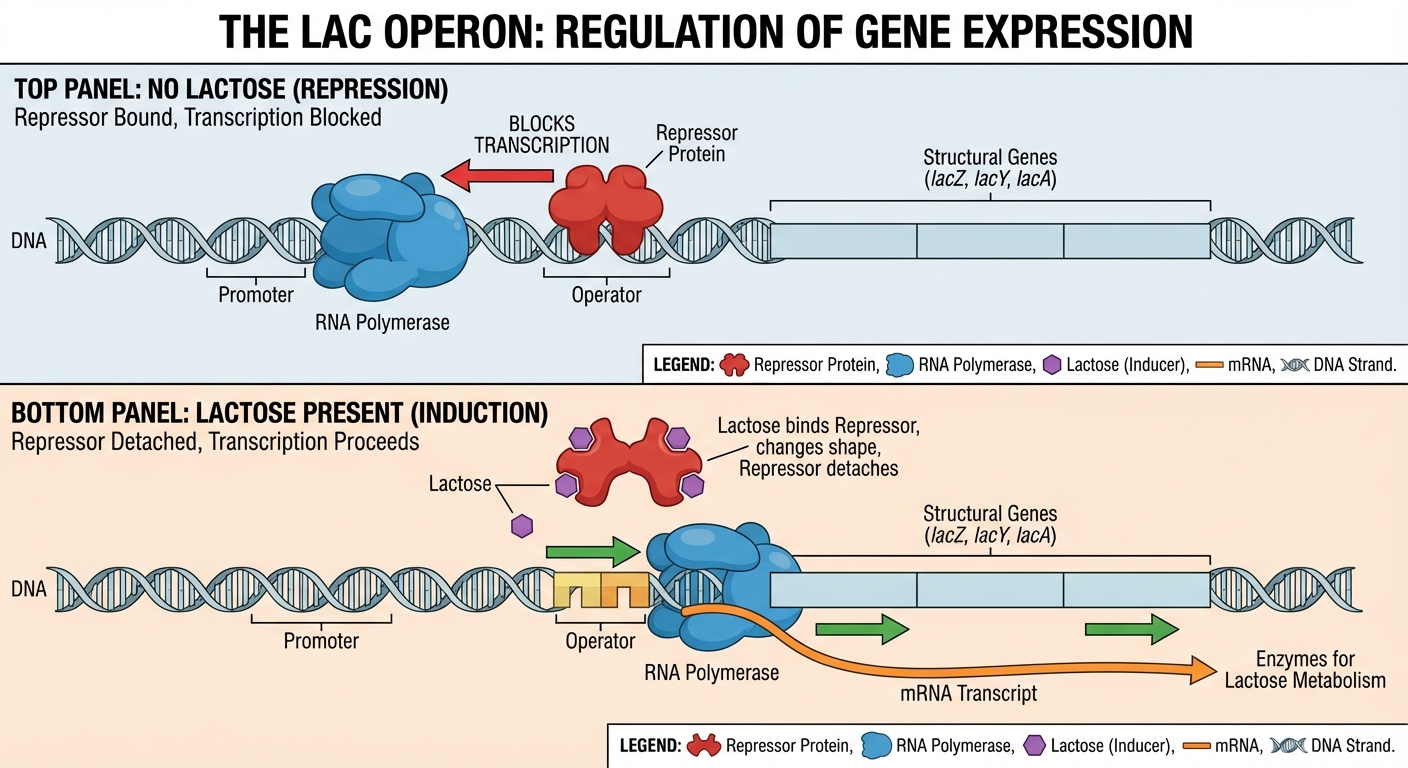
Inducible Operon (e.g., lac operon):
- Default State: OFF.
- Function: Catabolic (breaking things down).
- Mechanism: When Lactose is present (inducer), it binds to the repressor, changing its shape so it falls off the operator. Transcription turns ON.
Eukaryotic Regulation
Regulation is more complex and occurs at multiple levels:
- Epigenetics: Changes in gene expression without changing the DNA sequence.
- DNA Methylation: Adding methyl groups to DNA tightens it $\rightarrow$ Reduces transcription.
- Histone Acetylation: Adding acetyl groups helps DNA unwind $\rightarrow$ Increases transcription.
- Transcription Factors: Proteins that bind to specific DNA sequences (Enhancers or Silencers) to help RNA polymerase bind.
- Cell Specialization: Different cells express different transcription factors, leading to different tissue types (e.g., liver vs. skin cells) despite having the same genome.
- RNA Interference (RNAi): MicroRNAs (miRNAs) or siRNAs can bind to mRNA and degrade it or block translation (Post-transcriptional regulation).
Mutations
A mutation is a change in the genetic material of a cell.
Types of Point Mutations (Single Base Change)
- Silent Mutation: The base change creates a codon that codes for the same amino acid (due to redundancy). No effect.
- Missense Mutation: The base change codes for a different amino acid. Can range from no effect to major protein malfunction (e.g., Sickle Cell Anemia).
- Nonsense Mutation: The base change creates a STOP codon. Leads to a truncated (shortened), usually non-functional protein.
Frameshift Mutations
Caused by Insertions or Deletions of nucleotides (not in multiples of 3). This shifts the "reading frame," changing every amino acid downstream. These are usually disastrous.
Chromosomal Mutations
- Duplication: Extra copies of genes.
- Inversion: A chromosomal segment is reversed.
- Translocation: A segment breaks off and joins a non-homologous chromosome.
Biotechnology
Tools of the Trade
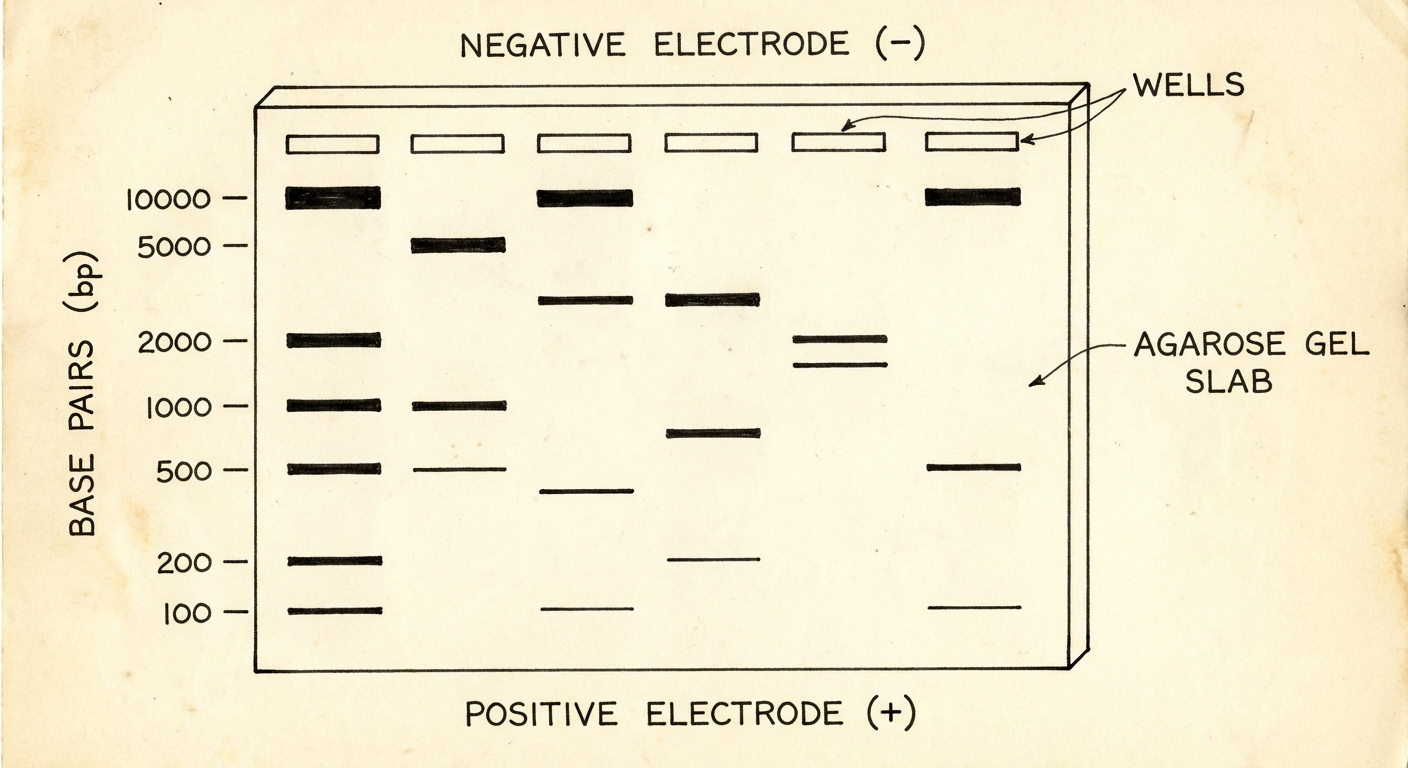
- Gel Electrophoresis: Separates DNA fragments by size and charge.
- DNA is negative (phosphate group), so it moves toward the positive electrode.
- Small fragments move faster; large fragments move slower.
- PCR (Polymerase Chain Reaction): Amplifies (copies) specific DNA segments billions of times rapidly. Uses heat-stable Taq polymerase.
- Transformation: Introducing foreign plasmid DNA into bacteria. Used to make bacteria produce insulin.
- DNA Sequencing: Determining the exact order of nucleotides in a genome.
- CRISPR-Cas9: A gene-editing tool derived from bacteria that allows scientists to cut and modify specific DNA sequences with high precision.
Common Mistakes in Unit 6
- Structure: Incorrectly stating that hydrogen bonds are in the backbone (they are between bases) or that covalent bonds are between bases (they are in the backbone).
- Replication vs. Transcription: Confusing DNA Polymerase (replication) with RNA Polymerase (transcription).
- Direction: Forgetting that new strands are always built . This means the template is read .
- Mutations: Assuming all mutations are harmful. Some are neutral, and some create the diversity necessary for evolution.