From Genotype to Phenotype: The Mechanisms of Gene Expression
1. The Central Dogma of Biology
Before diving into the mechanics, we must understand the fundamental framework of biology known as the Central Dogma. First proposed by Francis Crick, this concept explains the directional flow of genetic information within a biological system.
The typical flow is:
- Gene Expression is the process by which DNA directs the synthesis of proteins (or, in some cases, just RNAs like tRNA or rRNA).
- The sequence of bases in DNA determines the sequence of bases in RNA, which in turn determines the sequence of amino acids in a protein.
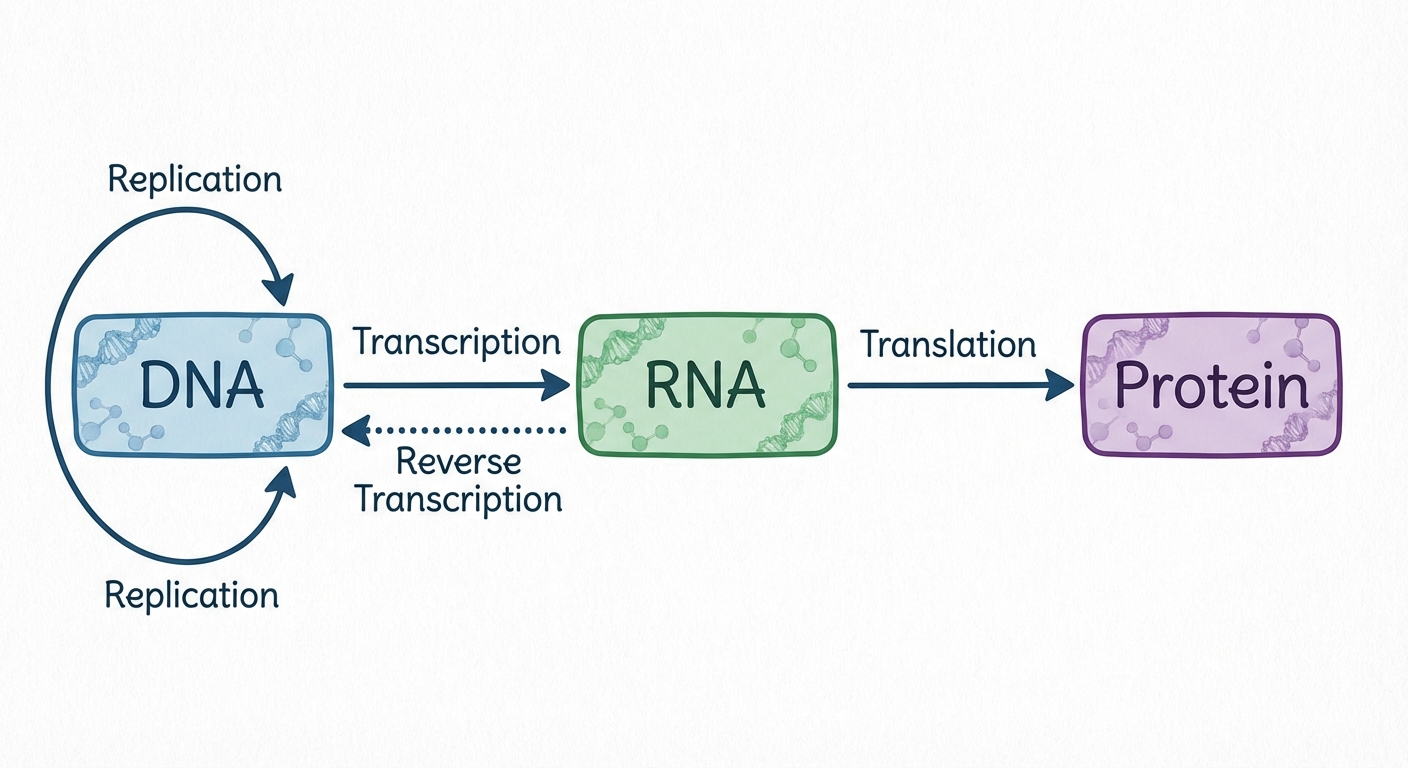
Exceptions to the Central Dogma
While the flow above is universal to cellular life, Retroviruses (like HIV) challenge this rule. They use an enzyme called Reverse Transcriptase to copy their RNA genome back into DNA (), allowing it to integrate into the host's genome.
2. Transcription: Synthesizing RNA
Transcription is the synthesis of an RNA molecule using information in DNA. This takes place in the nucleus of eukaryotic cells and the cytoplasm (nucleoid region) of prokaryotic cells.
The Key Players
- Template Strand (or Non-coding/Antisense strand): The DNA strand that is actually read by the enzyme. It is complementary to the RNA.
- Coding Strand (or Non-template/Sense strand): The DNA strand that is not read. Its sequence matches the RNA sequence exactly (except RNA has Uracil instead of Thymine).
- RNA Polymerase: The enzyme that pries the DNA strands apart and joins together RNA nucleotides complementary to the DNA template.
Directionality Rules
- RNA Polymerase adds nucleotides to the 3' end of the growing RNA molecule.
- Therefore, RNA is synthesized in the direction.
- The DNA template is read in the direction.
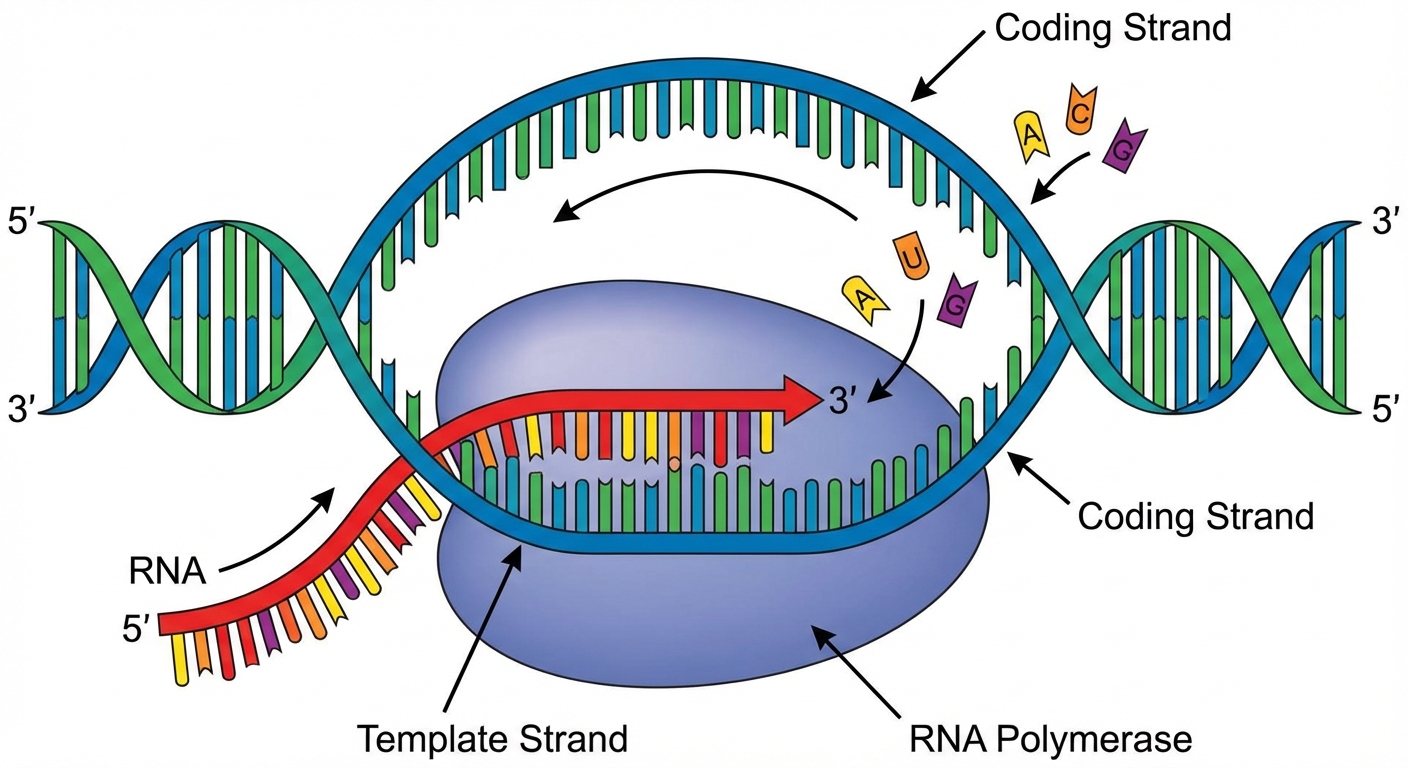
The Three Stages of Transcription
1. Initiation
- Promoter: A specific DNA sequence upstream of the gene where RNA Polymerase binds. In eukaryotes, the TATA box is a crucial promoter DNA sequence.
- Transcription Factors: In eukaryotes, these helper proteins must bind to the promoter before RNA Polymerase II can bind (forming the Transcription Initiation Complex).
2. Elongation
- RNA Polymerase moves along the DNA template, untwisting the double helix.
- It adds RNA nucleotides antiparallel and complementary to the template.
- DNA Adenine (A) $\to$ RNA Uracil (U)
- DNA Cytosine (C) $\to$ RNA Guanine (G)
- DNA Guanine (G) $\to$ RNA Cytosine (C)
- DNA Thymine (T) $\to$ RNA Adenine (A)
3. Termination
- Prokaryotes: Transcription stops at a specific terminator sequence; the RNA is released immediately.
- Eukaryotes: RNA Polymerase II transcribes a polyadenylation signal sequence (AAUAAA), which signals proteins to cut the RNA transcript free from the polymerase.
3. RNA Processing (Eukaryotes Only)
In prokaryotes, the RNA produced during transcription is ready to be translated immediately. In eukaryotes, the initial transcript (called pre-mRNA) must be modified in the nucleus before it is dispatched to the cytoplasm.
Steps of RNA Processing
- 5' GTP Cap: A modified Guanine nucleotide is added to the 5' end.
- Function: Protects from degradation and helps the ribosome attach.
- 3' Poly-A Tail: An enzyme adds 50–250 Adenine nucleotides to the 3' end.
- Function: Protects from degradation and assists in export from the nucleus.
- RNA Splicing: Removal of large portions of the RNA molecule.
- Introns (Intervening sequences): Non-coding segments that are cut out.
- Exons (Expressed sequences): Coding segments that are spliced (joined) together.
- Spliceosome: A large complex of proteins and small RNAs that executes the cutting and pasting.
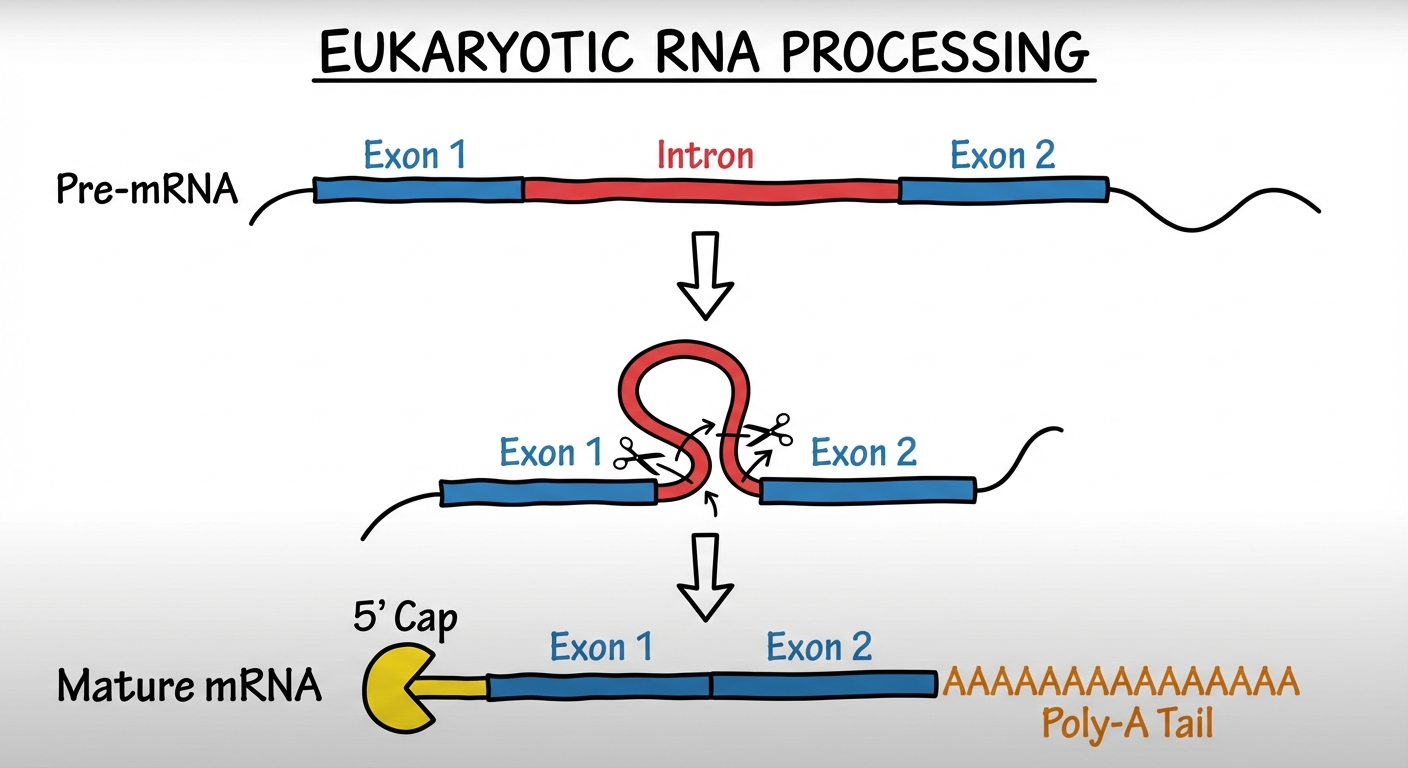
Alternative Splicing
This is a critical concept. A single gene can code for more than one kind of polypeptide, depending on which segments are treated as exons during splicing. This explains why humans have relatively few genes (~20,000) but can produce a vast array of proteins.
4. Translation: RNA to Protein
Translation is the RNA-directed synthesis of a polypeptide. It occurs on ribosomes in the cytoplasm (or on the Rough ER).
The Genetic Code
Genetic information is encoded as a sequence of non-overlapping base triplets called codons.
- Universality: The code is nearly universal (bacteria to humans use the same code), indicating common ancestry.
- Redundancy: There are 64 codons but only 20 amino acids. Most amino acids are specified by more than one codon (e.g., GGU, GGC, GGA, and GGG all code for Glycine).
- No Ambiguity: One specific codon always codes for the same amino acid (e.g., GGU is always Glycine).
Key Players
| Component | Function |
|---|---|
| mRNA | Carries the genetic message from DNA to the ribosome. |
| tRNA (Transfer RNA) | Transfers amino acids to the ribosome. Contains an Anticodon (complementary to mRNA codon). |
| rRNA (Ribosomal RNA) | Makes up the ribosome subunits; catalytically active (ribozyme) in peptide bond formation. |
The Ribosome Structure
The ribosome has three binding sites for tRNA:
- A site (Aminoacyl): Holds the tRNA carrying the next amino acid to be added.
- P site (Peptidyl): Holds the tRNA carrying the growing polypeptide chain.
- E site (Exit): Where discharged tRNAs leave the ribosome.
The Three Stages of Translation
1. Initiation
- The small ribosomal subunit binds to the mRNA and a specific initiator tRNA (carrying Methionine).
- It scans for the Start Codon (AUG).
- The large ribosomal subunit attaches, placing the initiator tRNA in the P site.
2. Elongation
Codon recognition $\to$ Peptide bond formation $\to$ Translocation.
- Recognition: The anticodon of an incoming tRNA pairs with the mRNA codon in the A site.
- Peptide Bond: An rRNA molecule catalyzes the formation of a peptide bond between the amino acid in the A site and the chain in the P site. The polypeptide is transferred to the A site tRNA.
- Translocation: The ribosome moves the tRNA from the A site to the P site. The empty tRNA in the P site moves to the E site and exits.
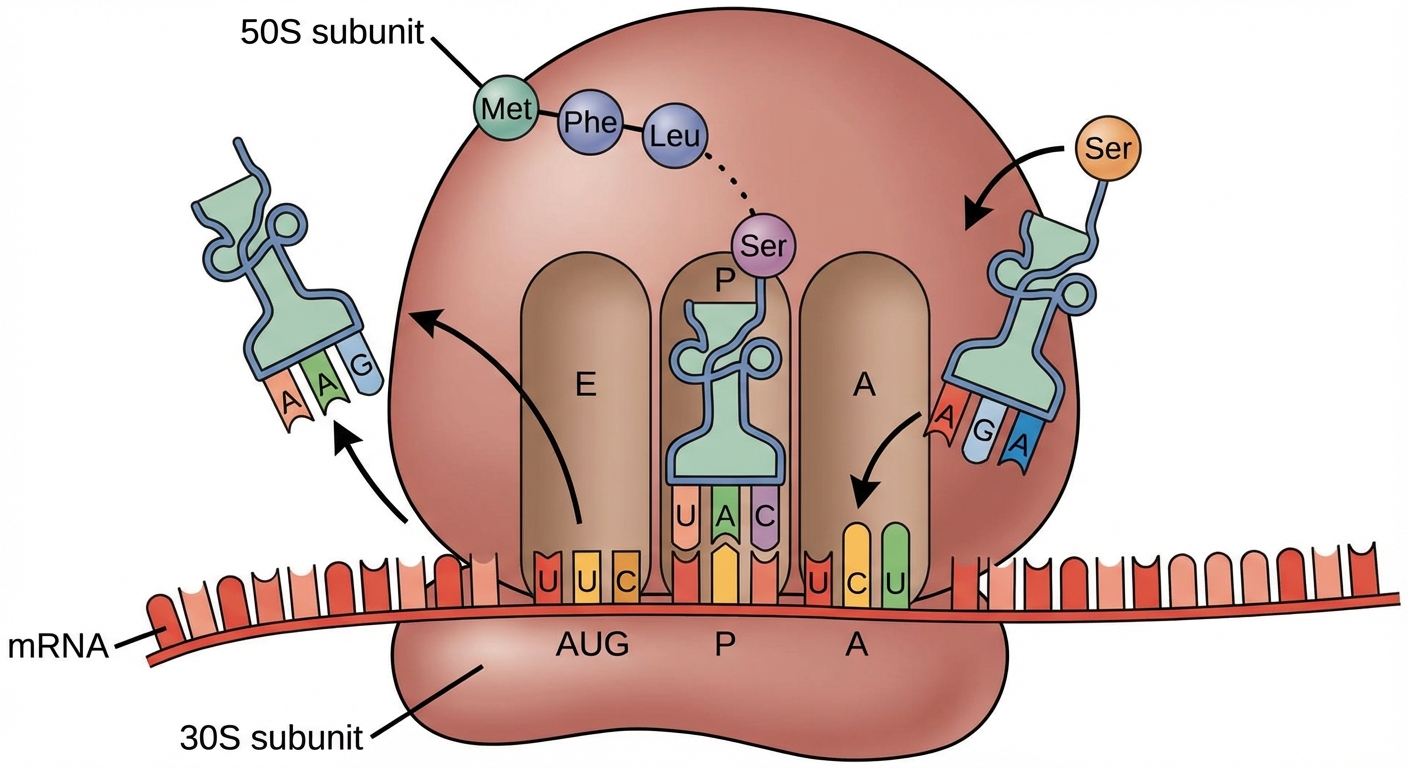
3. Termination
- Elongation continues until a Stop Codon (UAG, UAA, or UGA) reaches the A site.
- A Release Factor protein binds to the stop codon, adding a water molecule instead of an amino acid.
- This hydrolysis releases the polypeptide, and the translation assembly breaks apart.
5. Prokaryotes vs. Eukaryotes Summary
| Feature | Prokaryotes | Eukaryotes |
|---|---|---|
| Location | Transcription and Translation occur simultaneously in the cytoplasm. | Transcription in nucleus; Translation in cytoplasm. Separated by nuclear envelope. |
| RNA Processing | None. | Introns removed (splicing), 5' Cap, 3' Poly-A Tail. |
| Ribosomes | Smaller (70S). | Larger (80S). |
6. Common Mistakes & Pitfalls
- Confusing the DNA Strands: Students often try to transcribe the Coding strand. Remember: The Template strand is what RNA Pol reads. The resulting RNA looks like the Coding strand (U replacing T).
- Directionality Errors:
- Replication/Transcription: synthesize in (add to the 3' end).
- Ribosomes read mRNA in .
- However, the ribosome reads the DNA template (don't mix these up!).
- The Stop Codon: The stop codon is not an amino acid. It signals termination. Do not write "Stop" as part of the amino acid sequence.
- Mutations: Assuming a point mutation (change in one base) always changes the protein. Due to redundancy, a "silent mutation" might occur where the amino acid remains the same.
- Introns vs. Exons: A common mnemonic is "Exons are EXpressed (kept)" and "Introns go IN the trash."